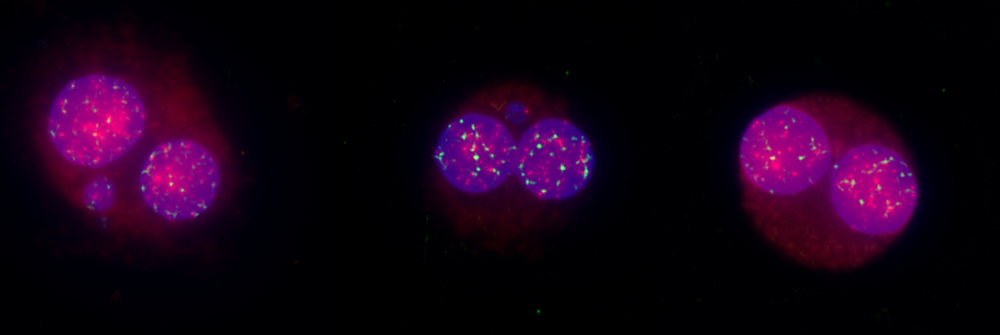
In vitro Micronucleus Test, Chromosomal Aberration Assay with Mode of Action
OECD TG487 – Principle of Micronucleus Test
When pharmaceutical, cosmetic, agrochemical, or chemical companies develop new compounds, predictive toxicology assays are essential. These typically begin with in vitro tests, which help determine whether in vivo testing is warranted. Among these assays, the in vitro micronucleus test stands out as one of the most successful and reliable methods for detecting chemicals with genotoxic potential.
Principle and Mechanism
Genotoxic chemicals or physical agents can cause two main types of genetic damage:
• Clastogenic events: Formation of acentric chromosome fragments
• Aneugenic events: Improper segregation of whole chromosomes
Both types of damage can result in the appearance of a micronucleus—a small, erratic nucleus separate from the main nucleus. The in vitro micronucleus test is specifically designed to detect clastogenic and aneugenic compounds, making it valuable for early screening and as part of regulatory test batteries.
Advantages and Application
• Ease of scoring: The assay is straightforward to interpret, and slide reading can be automated.
• Reliability: Structure-activity relationship (SAR) in silico models are less effective for predicting clastogenicity or aneugenicity, whereas FISH technology provides clear results.
• Versatility: The test is typically performed using human cell lines or human whole blood
In vivo micronucleus assays are also used, particularly on peripheral blood cells, for cytogenetic research and to investigate genotoxic effects of radiation.
Mechanistic Insights – Mode of Action
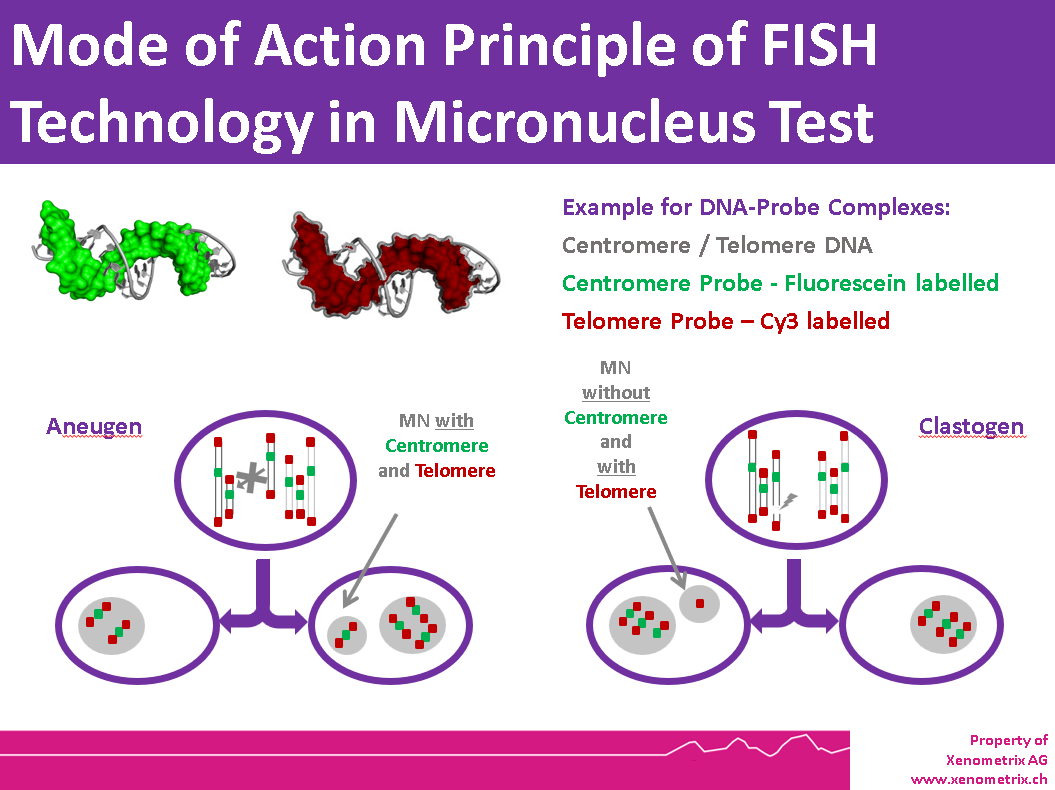
The characteristics of the micronucleus allow researchers to identify the mechanism of action, especially when FISH (fluorescence in situ hybridization) probes are applied. FISH probes allow the direct discrimination between clastogenic and aneugenic chemicals pharmaceuticals. This mechanistic information is crucial for understanding the biological effects of genotoxic agents and for designing follow-up studies.
OECD TG487 Protocol Overview
The micronucleus assay is described in OECD Test Guideline 487. Key steps include:
• Mammalian cells are cultured and treated with the test compound during exponential growth.
• Cells must undergo at least 1.5–2 cell cycles during exposure.
• Treatment schedules vary depending on cell type (cell lines, primary cultures, or lymphocytes).
• Most aneugens and clastogens are detectable after 3–6 hours of treatment, with or without metabolic activation.
• After exposure, the test compound and S9 are removed, fresh media is added, and cells are incubated for 2 cell cycles.
Cells are then harvested and prepared for slide analysis (hypotonic treatment or cell spreading). Micronuclei can be stained using various techniques, such as DAPI or FISH probes for centromere and telomere detection. Fluorescent dyes help eliminate artifacts from non-DNA-specific stains. FISH with pancentromeric and telomeric DNA probes, combined with DAPI counterstaining, enables identification of micronucleus contents and provides information on the mode of action.
Comparison to Chromosomal Aberration Test
The in vitro micronucleus test is generally preferred over the chromosomal aberration test because it can detect chromosome loss (aneugenicity), which the chromosomal aberration test cannot.
Micronucleus Test with FISH Probes - Rapid MicroNucleus MoA
FISH probes and a set of carefully optimized solutions allow to detect centromeres and telomeres in human cells sensitively, using both ex vivo human full blood or human derived cell lines, in as fast as 3 hours. A ready to use kit system is available: Rapid Micronucleus MoA Centromere Telomere.
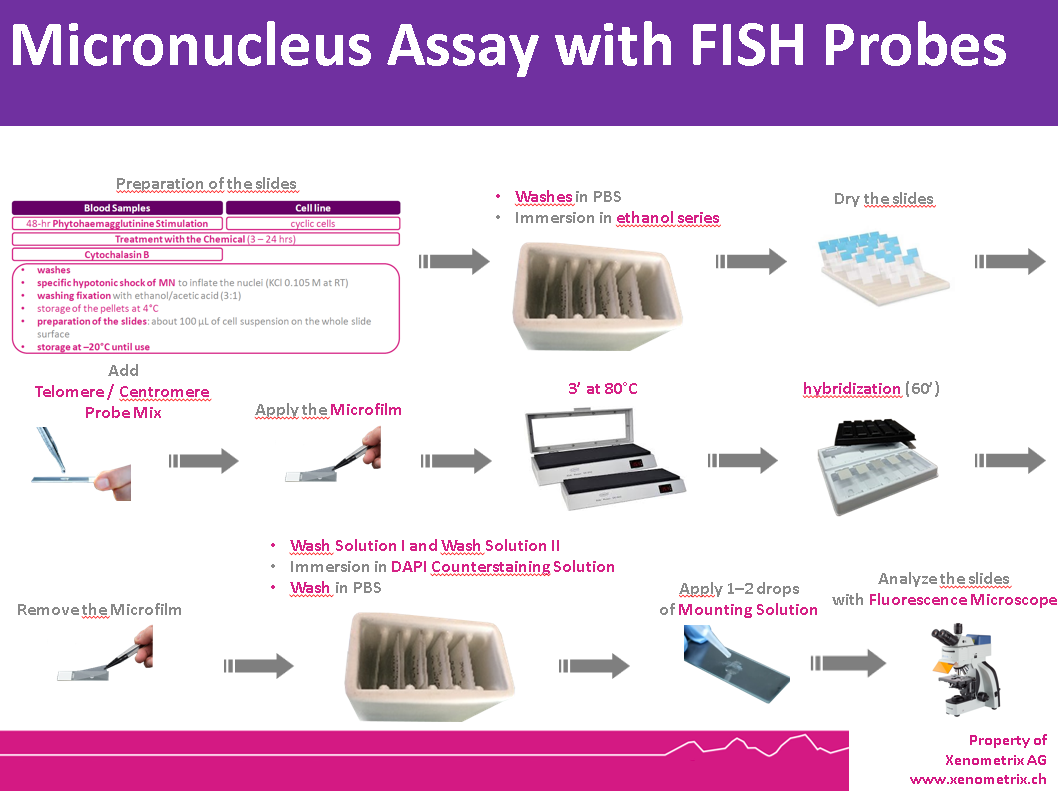
Cells are prepared as per standard protocols according to OECD TG 487 and the slides are sequentially treated with a series of solutions provided with the kit “Rapid Micronucleus MoA”, including human-specific, highly sensitive centromere and telomere probes and a counterstaining solution containing DAPI (4’,6-diamidino-2-phenylindole). In detail, slides are sequentially washed with PBS and an ethanol series after treatment. Centromere and Telomere probes are hybridized with the DNA of the sample at 80°C and transferred to a humidified chamber. Washes and a counterstaining step follow prior to the addition of the mounting solution.
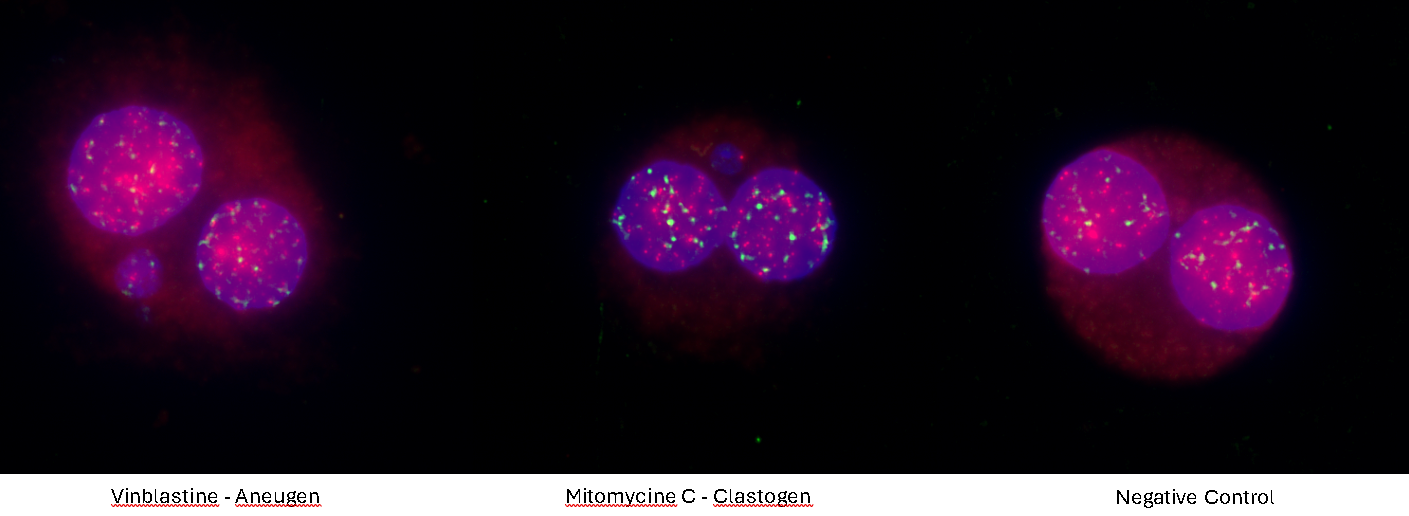
Using a 40× magnification objective it will be possible to easily identify micronuclei and assess whether a clastogenic or aneugenic event has happened. Automated reading and archiving can be obtained with slide scanning systems, e.g. Metafer. Slides are usually ready for evaluation within 1 hour.
Biomonitoring the exposition to low doses of radiation - FISH‑based micronucleus‑centromere assay
FISH probe technology as used in the Rapid Micronucleus MoA Kit can also be used in cytogenetic analysis, in addition to physical dosimetry, i.e. people exposed in medical or natural radiation.
Fast, Sensitive Detection of Mode of Action with Micronucleus Assay
The Xenometrix innovative kit combines advanced FISH probe technology with carefully optimized solutions to detect aneugenic and clastogenic compounds - delivering results in as little as 3 hours. Whether using ex vivo human whole blood or human-derived cell lines, the Rapid Micronucleus MoA Centromere Telomere Kit delivers reliable genotoxicity screening.
Key Features:
• Ready-to-Use System: All reagents and probes are included for seamless integration into your lab.
• OECD TG 487 Compliant: Standardized protocols ensure regulatory acceptance and scientific rigor.
• Human-Specific, Highly Sensitive Probes: Detect centromeres and telomeres with exceptional clarity.
• Quick Turnaround: Slides are ready for evaluation within 3 hours after preparation.
How It Works:
1. Cells are prepared following OECD TG 487 or OECD TG473 guidelines.
2. Slides are treated with a series of kit solutions, including centromere and telomere FISH probes and DAPI counterstain.
3. Probes are hybridized at 80°C, followed by washing and mounting.
4. Using a 40× magnification objective, micronuclei are easily identified and clastogenic and aneugenic events are visible.
5. Automated slide scanning and archiving (e.g., Metafer systems) enable efficient, high-throughput analysis.
Applications:
• Genotoxicity Screening: Rapidly assess chromosomal damage in research and product development.
• Biomonitoring Radiation Exposure: Sensitive detection of low-dose radiation effects in medical or environmental settings.
More information.
Rapid MicroNucleus MoA – References
- OECD (2023), Test No. 487: In Vitro Mammalian Cell Micronucleus Test, OECD Guidelines for the Testing of Chemicals, Section 4, OECD Publishing, Paris, https://doi.org/10.1787/9789264264861-en.
- Cuceu Petrenci, C., Raposo, J., Lacour, T., & Mouche, I. (2025). P30-59 Interest of the Micronucleus test on Reconstructed Skin and skin organoids for cosmetic products. Toxicology Letters, 411(Suppl.), S398–S399. https://doi.org/10.1016/j.toxlet.2025.07.918
- OECD (2016), Test No. 473: In Vitro Mammalian Chromosomal Aberration Test, OECD Guidelines for the Testing of Chemicals, Section 4, OECD Publishing, Paris, https://doi.org/10.1787/9789264264649-en.
- www.metasystems-international.com
- Maja Kneissig, Kristina Keuper, Mirjam S de Pagter, Markus J van Roosmalen, Jana Martin, Hannah Otto, Verena Passerini, Aline Campos Sparr, Ivo Renkens, Fenna Kropveld, Anand Vasudevan, Jason M Sheltzer, Wigard P Kloosterman, Zuzana Storchova (2019) Micronuclei-based model system reveals functional consequences of chromothripsis in human cells eLife 8:e50292
- Schupp N, Stopper H, Heidland A. DNA Damage in Chronic Kidney Disease: Evaluation of Clinical Biomarkers. Oxid Med Cell Longev. 2016;2016:3592042. doi: 10.1155/2016/3592042. Epub 2016 May 25. PMID: 27313827; PMCID: PMC4897719.
- Ladeira C, Smajdova L. The use of genotoxicity biomarkers in molecular epidemiology: applications in environmental, occupational and dietary studies. AIMS Genet. 2017 Aug 11;4(3):166-191. doi: 10.3934/genet.2017.3.166. PMID: 31435507; PMCID: PMC6690241.
- Takeiri, A., Matsuzaki, K., Motoyama, S. et al. High-content imaging analyses of γH2AX-foci and micronuclei in TK6 cells elucidated genotoxicity of chemicals and their clastogenic/aneugenic mode of action. Genes and Environ 41, 4 (2019). https://doi.org/10.1186/s41021-019-0117-8
- Natarajan A.T., Boei J.J., Darroudi F., Van Diemen P.C., Dulout F., Hande M.P. and Ramalho A.T. 1996. Current cytogenetic methods for detecting exposure and effects of mutagens and carcinogens. Environ Health Perspect 104(Suppl 3):445–448
- Thierens H., Vral A., Morthier R., Aousalah B. and De Ridder L. 2000. Cytogenetic monitoring of hospital workers occupationally exposed to ionizing radiation using the micronucleus centromere assay. Mutagenesis 15:245–249
- Fenech M., Kirsch-Volders M., Natarajan A.T., Surralles J., Crott J.W., Parry J., Norppa H., Eastmond D.A., Tucker J.D. and Thomas P. 2011. Molecular mechanisms of micronucleus, nucleoplasmic bridge and nuclear bud formation in mammalian and human cells. Mutagenesis. 26(1):125–132
- Vral A., Decorte V., Depuydt J., Wambersie A., Thierens H. 2016. A semiautomated FISH based micronucleuscentromere assay for biomonitoring of hospital workers exposed to low doses of ionizing radiation. Molecular medicine reports 14:103–10

 Navigation
Navigation